Prediction of a fractional SPDE using the covariance-based rational SPDE approximation
Source:R/fractional.computations.R
predict.CBrSPDEobj.RdThe function is used for computing kriging predictions based on data \(Y_i = u(s_i) + \epsilon_i\), where \(\epsilon\) is mean-zero Gaussian measurement noise and \(u(s)\) is defined by a fractional SPDE \((\kappa^2 I - \Delta)^{\alpha/2} (\tau u(s)) = W\), where \(W\) is Gaussian white noise and \(\alpha = \nu + d/2\), where \(d\) is the dimension of the domain.
Arguments
- object
The covariance-based rational SPDE approximation, computed using
matern.operators()- A
A matrix linking the measurement locations to the basis of the FEM approximation of the latent model.
- Aprd
A matrix linking the prediction locations to the basis of the FEM approximation of the latent model.
- Y
A vector with the observed data, can also be a matrix where the columns are observations of independent replicates of \(u\).
- sigma.e
The standard deviation of the Gaussian measurement noise. Put to zero if the model does not have measurement noise.
- mu
Expectation vector of the latent field (default = 0).
- compute.variances
Set to also TRUE to compute the kriging variances.
- posterior_samples
If
TRUE, posterior samples will be returned.- n_samples
Number of samples to be returned. Will only be used if
samplingisTRUE.- only_latent
Should the posterior samples be only given to the laten model?
- compute_var_method
Method to compute the variances. Options are: "direct", "loop", and "selected_inv". The "direct" method computes the variance directly, which is faster but can be memory intensive. The "loop" method computes the variance by looping over the elements, which is more memory efficient but slower.
- ...
further arguments passed to or from other methods.
Value
A list with elements
- mean
The kriging predictor (the posterior mean of u|Y).
- variance
The posterior variances (if computed).
Examples
set.seed(123)
# Sample a Gaussian Matern process on R using a rational approximation
kappa <- 10
sigma <- 1
nu <- 0.8
sigma.e <- 0.3
range <- 0.2
# create mass and stiffness matrices for a FEM discretization
x <- seq(from = 0, to = 1, length.out = 101)
fem <- rSPDE.fem1d(x)
tau <- sqrt(gamma(nu) / (sigma^2 * kappa^(2 * nu) *
(4 * pi)^(1 / 2) * gamma(nu + 1 / 2)))
# Compute the covariance-based rational approximation
op_cov <- matern.operators(
loc_mesh = x, nu = nu,
range = range, sigma = sigma, d = 1, m = 2,
parameterization = "matern"
)
# Sample the model
u <- simulate(op_cov)
# Create some data
obs.loc <- runif(n = 10, min = 0, max = 1)
A <- rSPDE.A1d(x, obs.loc)
Y <- as.vector(A %*% u + sigma.e * rnorm(10))
# compute kriging predictions at the FEM grid
A.krig <- rSPDE.A1d(x, x)
u.krig <- predict(op_cov,
A = A, Aprd = A.krig, Y = Y, sigma.e = sigma.e,
compute.variances = TRUE
)
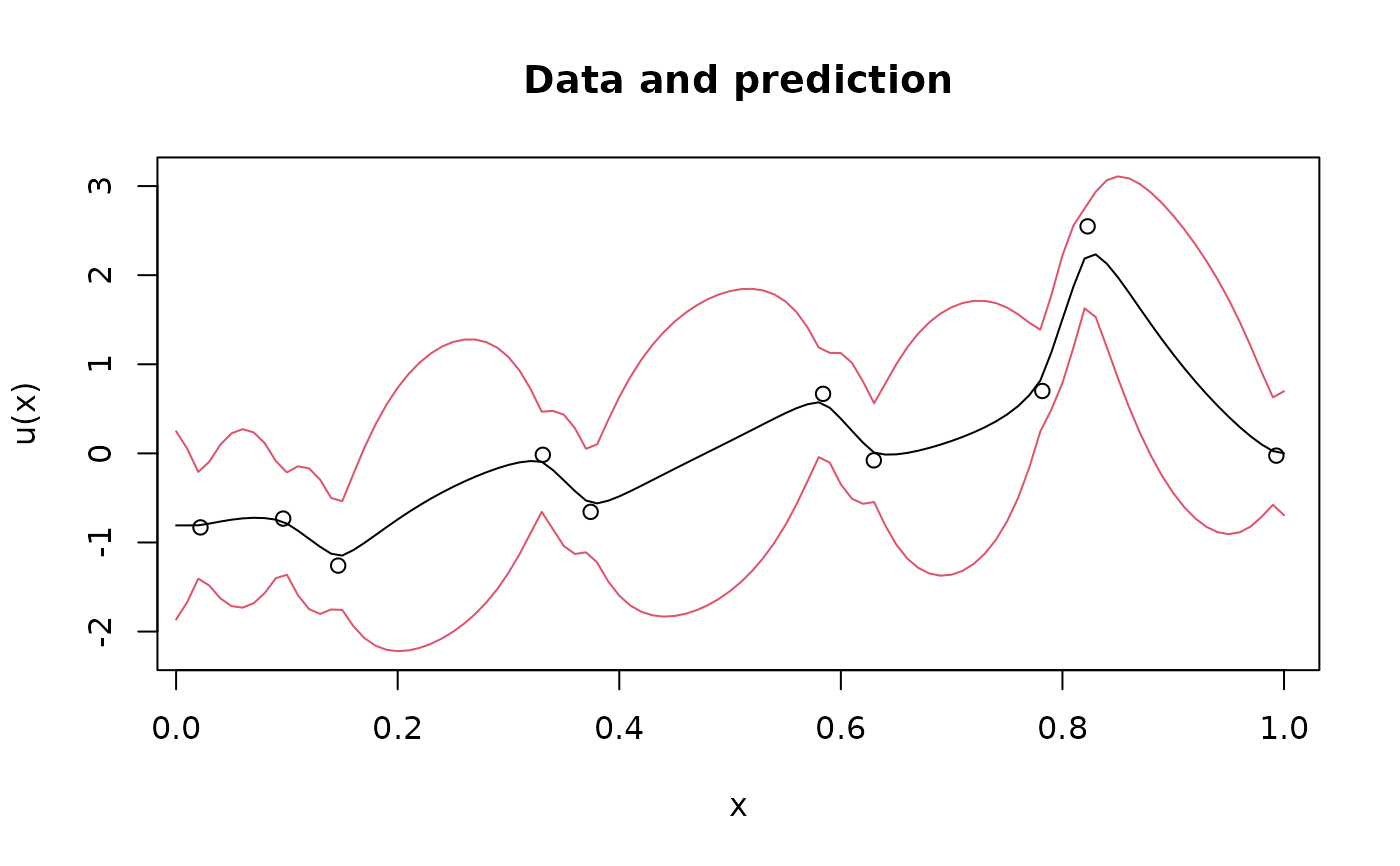
plot(obs.loc, Y,
ylab = "u(x)", xlab = "x", main = "Data and prediction",
ylim = c(
min(u.krig$mean - 2 * sqrt(u.krig$variance)),
max(u.krig$mean + 2 * sqrt(u.krig$variance))
)
)
lines(x, u.krig$mean)
lines(x, u.krig$mean + 2 * sqrt(u.krig$variance), col = 2)
lines(x, u.krig$mean - 2 * sqrt(u.krig$variance), col = 2)